GauSwastha —
Cattle Health AI
A camera-only cattle health and valuation engine that estimates weight, body condition score, breed grade, and conformation traits from a single smartphone photo — deployed with the Government of Karnataka across 30+ polyclinics.
Cattle assessment is broken for smallholders
India has over 300 million cattle, the majority owned by smallholder farmers with 2–10 animals. For these farmers, knowing the health, weight, and breed quality of their livestock isn't just agricultural knowledge — it determines loan eligibility, insurance valuation, and the price they get at market.
Yet the current state of assessment is either manual — tape measurements, visual inspection, and experience-based guesses that vary wildly between vets — or hardware-driven, using LiDAR scanners and depth cameras that cost lakhs and exist only in research labs or large commercial farms.
The smallholder with five cows in a village shed has no access to either. When a government vet visits, they spend a few minutes per animal and move on. There is no objective, repeatable record. No baseline to track health over time. No way to dispute a low insurance payout.
Can we recover enough geometric and visual information from a single ordinary smartphone photo to approximate what sensor-based systems need specialised hardware to measure — and deliver it in under two seconds on a CPU server in a rural clinic?

What existing research gets wrong
Most academic work on automated cattle monitoring assumes access to specialised hardware: LiDAR scanners, structured-light depth cameras, walk-over weigh bridges, or fixed multi-camera rigs in large commercial barns. These deliver high-quality 3D point clouds and achieve strong accuracy, but are capital-intensive and evaluated on organised farms with hundreds of animals in controlled conditions.
My challenge was to reframe the same goals — weight, body condition score, conformation — for the reality of small, unorganised farmers who own at most a handful of cows and whose only guaranteed device is a mid-range Android phone.
Most papers rely on LiDAR or depth cameras positioned around the animal. These capture accurate 3D data but are expensive, fragile, and impossible to maintain outside controlled research barns or large commercial farms.
A common assumption is a fixed camera position or raceway where every animal walks through the same calibrated zone. This doesn't match open yards, village sheds, or the ad-hoc spaces where smallholders actually handle their cattle.
Academic literature largely targets large commercial farms that can justify high capital expenditure and dedicated technical staff. Smallholders with five to ten cows cannot invest in fixed rigs or wearables — and almost none of the proposed systems translate into something they could realistically buy or use.
There was no published work on deriving veterinary-grade cattle metrics from a single handheld smartphone photo taken in uncontrolled field conditions by a non-expert. This became the design constraint for everything that followed.
Building a dataset in the field, not the lab
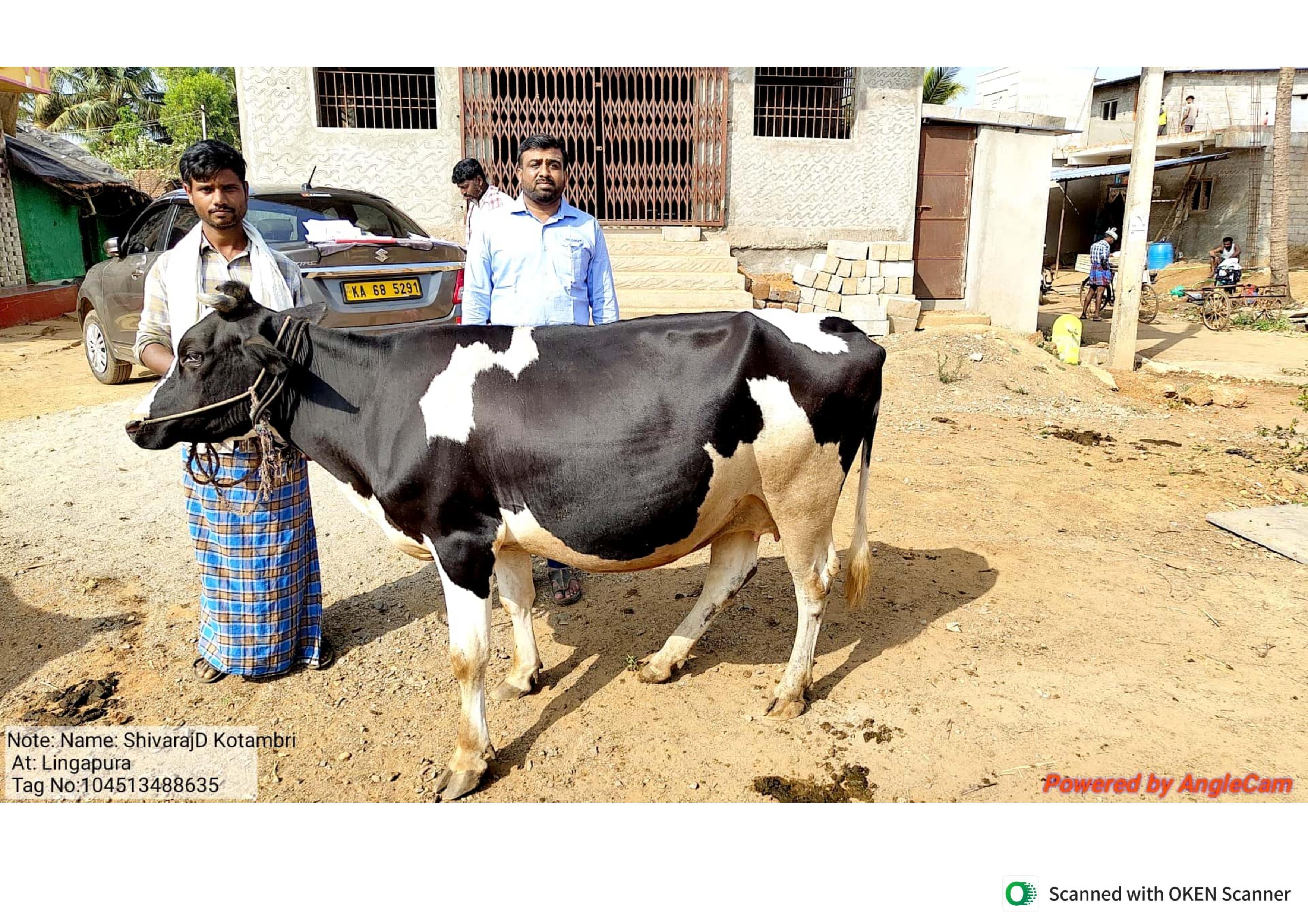
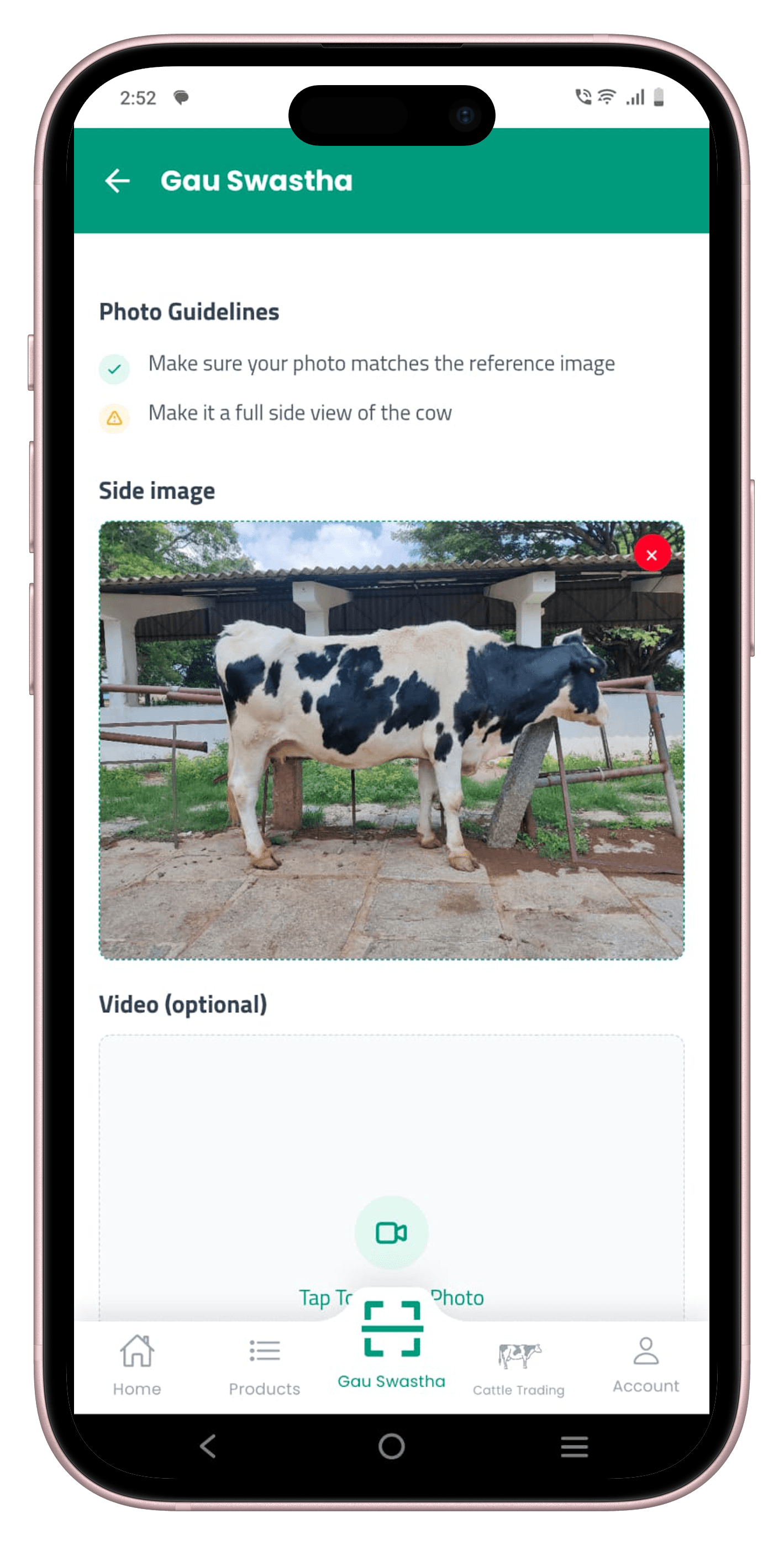
Cattle images were captured on mid-range Android phones by government vets and field staff during routine examinations across Karnataka. There was no fixed rig or controlled walkway — animals were photographed in sheds, open yards, and roadside spots at arm's length, however the farmer was keeping the animal still at that moment.
This keeps the data honest to how the system will actually be used. But it means the dataset is noisy, cluttered, and highly variable in lighting, background, animal pose, and distance — exactly the kind of distribution shift that kills lab-trained models in production. We treated this as a feature, not a bug.
We started by capturing front, rear, and side views of each animal. After running early experiments, the side view consistently gave the best trade-off between field practicality and predictive accuracy — it captures the full dorsal profile, rib visibility, and udder, which drive the most important downstream estimates. We standardised the dataset to a single left-side view.
~3,000 images collected across Karnataka, covering front, rear, and side views before standardisation to a single side view. 72 class labels covering body parts, visual traits, and six priority disease markers. Each image labelled in under a minute by trained veterinary staff.
Training vets to be labellers
The bottleneck in any medical or veterinary ML project isn't data — it's labelled data. General-purpose annotation services can draw bounding boxes but can't reliably distinguish a BCS-2.25 cow from a BCS-3 one, or identify early-stage udder disease. That knowledge lives with the vets in the field.
I designed a lightweight annotation protocol anchored in clinical vocabulary: regions vets already reason about — head, neck, torso, ribs, legs, udder — became the class taxonomy. I ran short onboarding sessions to train vets to annotate directly on their phones using Roboflow's mobile interface. For each image, they drew bounding boxes around key regions and assigned one of 72 class labels to every box.
The protocol balanced two constraints: annotations had to be quick enough to fit into a busy clinic workflow (target: under a minute per image), yet structured enough to drive downstream computer vision models. The 72-class label universe was deliberately broad — it made the dataset future-proof, letting us re-group the same labels into different targets as we experimented with new modelling approaches without needing to re-annotate.
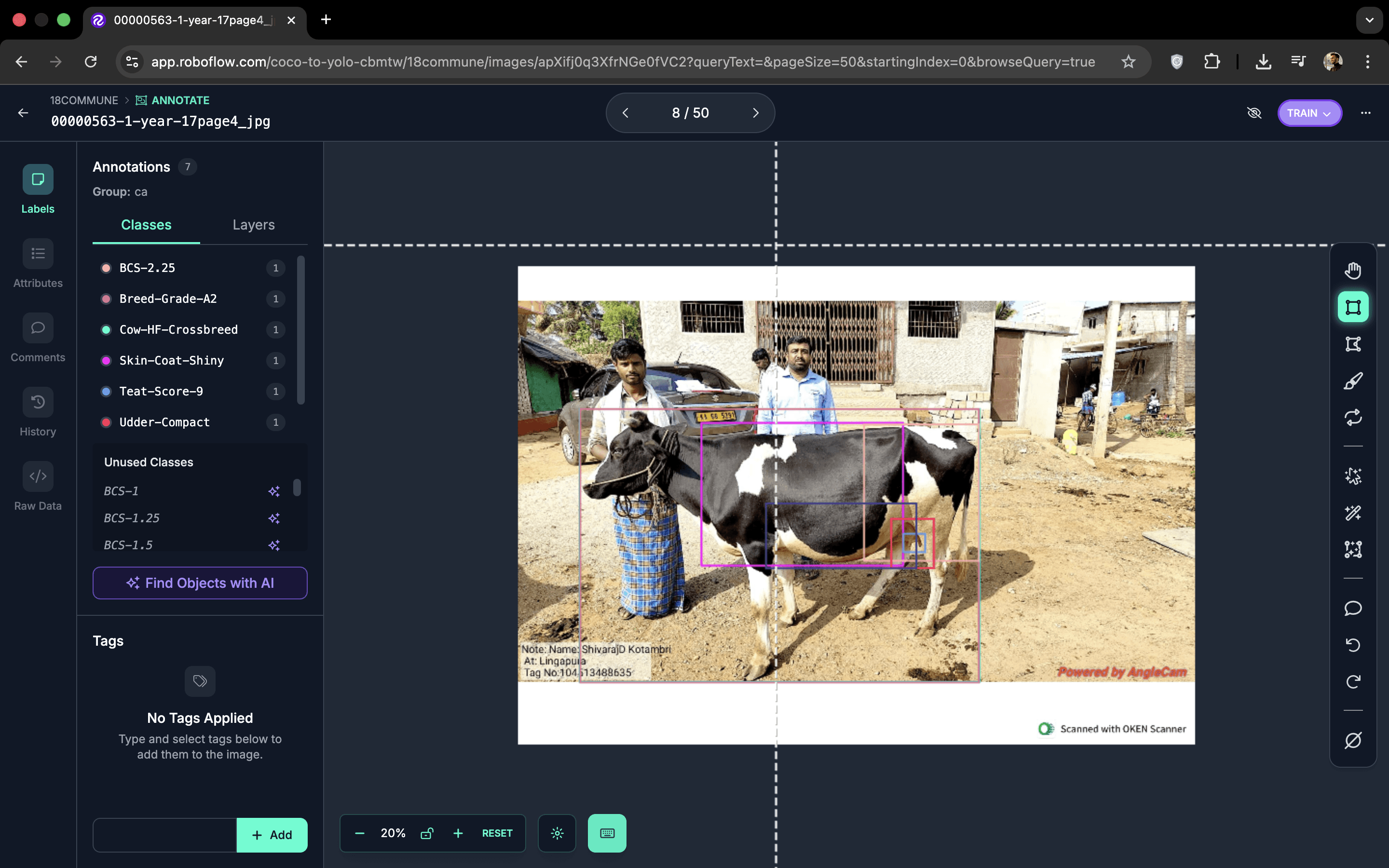
Body parts, visual traits (rib visibility, udder fullness, coat quality), and six priority disease markers. Made the dataset future-proof — the same annotated images drove all model heads without re-labelling as objectives evolved.
Vets labelled images in short bursts between real consultations. The annotation workflow was designed to feel like clinical note-taking, not a separate technical task — which kept the label quality high and the vets willing to do it.
From a single photo to a vet-style report
The system runs a multi-stage computer vision and ensemble pipeline, designed to work on CPU-only infrastructure in rural government clinics. Every stage is independently testable and replaceable — important in a domain where the data distribution shifts as we expand to new geographies and cattle breeds.
Standardise orientation, crop to the side-view bounding box, resize to inference resolution. Handles wide variation in phone cameras, focal lengths, and lighting conditions across thousands of field submissions.
Detects anatomical landmarks — hip pins, rump, spine, ribs, udder attachment — and returns bounding boxes and keypoints used by all downstream regression and classification heads.
Separate regression heads for live weight (kg) and body condition score (1–5 scale). Each head is trained on the detected anatomical regions from step 2, not on the full image — this is what makes the estimates robust to background clutter.
Breed classification, coat quality assessment, posture scoring, and milk yield estimation. A majority-vote ensemble across multiple crop windows reduces single-frame noise — critical when the vet captures the animal mid-step or at a slight angle.
All model outputs are aggregated into a structured veterinary-style report: breed grade, BCS, approximate weight, milk production capacity, lactation limit, and economic parameters. Served via FastAPI. Average latency ~1.2s on CPU. Delivered via the GauSampurna app and WhatsApp.
Steps 2–4 are powered by 24 independent YOLO models arranged in three layers — each specialised for one prediction task, with outputs from earlier layers feeding directly into later ones. The full architecture is detailed in the next section.
24 specialized models, one cascading pipeline
The naive approach would be a single model with many heads. We tried it. It underperformed on minority classes and was brittle when the input distribution shifted across breeds and geographies. The solution was a modular cascade: 24 independent YOLO models, each responsible for one specific prediction task, arranged in three layers.
Outputs from earlier layers become inputs to later ones. A body-bounds crop from Layer 1 is the input image for every Layer 2 classifier. The BCS-head, BCS-torso, and BCS-hindquarters models each see a different anatomical crop; their probability distributions are averaged by an ensemble in Layer 3 to produce the final BCS score. Breed + breed-grade then feed the rules engine, which uses them alongside teat score and udder type to calculate milk yield range and economic value — outputs no single model ever directly predicts.
This architecture meant each module could be developed, evaluated, and improved in isolation. When the worm-load classifier underperformed on a particular breed, we fixed that one module without touching anything else in the cascade.
Detect
Engine
Across all 24 modules in production, the system averaged 92% accuracy predicting the right class. BCS was the hardest — a continuous score binned into fine-grained increments that vets themselves disagree on. The three-crop ensemble (head, torso, hindquarters) consistently outperformed any single BCS model by 4–6 percentage points.
BCS ensemble. Body Condition Score uses a 1–5 scale with half and quarter increments. Three YOLO classifiers each see a different anatomical crop — head region, mid-torso, and hindquarters. Their class probability distributions are averaged and the final BCS is the argmax of the fused distribution. The ensemble consistently reduced variance across different capture angles and outperformed any single-crop model on every holdout set we evaluated.
Weight hybrid. Live weight is estimated through a hybrid approach combining monocular depth cues, keypoint-derived geometric proxies, and a trained regressor. The derivation and ablation are described in the BosWeigh experiment writeup.
The hardest part of production CV: preventing garbage in
In a lab setting every image you evaluate is a cattle photo taken under reasonable conditions. In production, users upload blurry shots, images of dogs, selfies, and unrelated field scenes. Without a gating layer, every such input silently passes through 24 models and produces a nonsense report — or worse, a plausible-looking one with confident-sounding numbers built on nothing.
The gates are the first three modules in Layer 1: cattle-gate (is there a cow?), view-gate (is it a left-side view?), and quality-gate (is the image sharp and well-lit enough?). They run sequentially — a failed gate short-circuits the pipeline immediately, invoking no downstream model.
A false negative from any gate — a real, valid cattle photo that gets rejected — means every downstream module receives no input and the user receives no report. In a paid product running 500+ scans daily, even a 2% gate false-negative rate means ten customers per day seeing a blank result they paid for. That is the fastest path to churn, and it compounds: a user who gets a null result once rarely tries again.
Trained to distinguish cattle from all other inputs. Threshold calibrated for high recall — we would rather pass a marginal cattle photo and let downstream models degrade gracefully than silently reject a valid paid scan. False positives here are recoverable; false negatives are not.
Validates a left-side profile view. Front and rear shots break every geometric assumption the downstream pipeline relies on, producing catastrophic failures across all Layer 2 and 3 modules. This gate has zero tolerance for view mismatch.
Checks blur threshold, minimum resolution, and lighting sufficiency. A blurry udder region makes teat-score predictions meaningless; heavy backlight breaks coat quality and BCS assessments. Gate failures return specific, actionable error messages rather than a generic reject.
Each gate failure returns a specific prompt: "Step back 1.5m", "Move into better light", "Ensure the full body is visible". A specific instruction the user can act on caused far lower churn than a silent failure or generic error. This was validated empirically through support ticket rate before and after the change.
Gate threshold calibration was the most iterative part of the build. Set too strict and too many real scans are rejected. Too lenient and bad inputs cascade into downstream partial failures — where some modules return outputs and others do not, leaving users unable to tell which parts of the report to trust. The final configuration optimised for recall at the gate level and let downstream modules handle ambiguity through confidence scores and low-confidence flags rather than null outputs.
95% cost reduction with serverless inference
Running 24 models in sequence, even on CPU, adds up. The first production setup was a FastAPI server on an always-on EC2 instance — simple, reliable, and expensive. At 500 scans per day concentrated in clinic hours, the instance was idle more than 95% of the time, billing compute budget around the clock for doing nothing.
The solution was AWS Lambda — pay per invocation, not per hour. Lambda executes the inference function on request, keeps the execution environment warm between calls, and scales to zero when idle. For a usage pattern with clear daytime peaks and long overnight troughs, this was a near-perfect operational fit.
vs. always-on EC2. Lambda's pay-per-request model eliminated idle-time billing across nights and weekends entirely.
End-to-end on a warm Lambda instance — all 24 models, gates included, report generated and returned.
Fully CPU-based inference. No GPU instances, no driver maintenance, no cold-GPU spin-up latency.
YOLO models are stored in S3 and downloaded to Lambda's /tmp directory on cold start. On warm invocations the execution context persists — models stay resident in memory between requests. This eliminated per-request model-load overhead that made early attempts too slow to be viable.
Cold starts add 3–5s when a new Lambda instance initialises from scratch. Provisioned concurrency kept warm instances ready during clinic hours. Off-peak cold starts were acceptable — users understood occasional delays outside business hours and the cost savings more than justified the trade-off.
Layer 1 gates run strictly sequentially — each must pass before the next fires. Layer 2 classifiers run in parallel threads: breed and udder type execute simultaneously, disease markers run in a separate thread pool. Layer 3 waits for all Layer 2 outputs before ensembling. This cut total inference time by ~40% vs naive sequential execution of all 24 models.
Not every user installs the GauSampurna app. Running the same Lambda pipeline behind a WhatsApp bot widened distribution to vets with older phones and farmers unwilling to install another app. Same backend, same 24 models, same report format — just a different input/output channel.
The full breakdown — model packaging, container vs zip deployment, memory tuning, and exact cost comparison before and after — is on Medium: How we cut AI inference costs by 95% using AWS Lambda →
What we learned from experimentation
The path from first prototype to production involved a series of decisions that weren't obvious upfront. Each one simplified the system without sacrificing accuracy — which mattered because the system had to be maintainable by a small team with limited compute.
We started collecting front, rear, and side images. During modelling, the side view alone gave the best accuracy while cutting the data collection burden by two-thirds. Simplifying to one view also made the annotation faster and the field protocol clearer for vets.
A deliberate constraint from day one — rural clinics don't have GPU servers, and we wanted per-scan costs low enough to sustain a paid product. This forced model compression, quantisation, and efficient head design. We achieved under 1.5s latency on a standard cloud CPU instance.
Instead of fixing the prediction target early, the 72-class label universe supported both object detection and multiple downstream tasks. The same images drove all model heads. When we added milk yield estimation months later, we didn't need to re-label a single photo.
Domain labels like BCS and breed grade require genuine veterinary knowledge. Outsourcing to a general annotation service would have introduced systematic errors in the most important targets. Vet-annotated data consistently produced better downstream models on every metric we tracked.
Production impact
GauSwastha went from a research prototype to a government-deployed production system. The GauSampurna Android app, which wraps the model, reached 100,000 downloads organically through vet networks across Karnataka — without any paid marketing. A paid pilot running at 30+ government polyclinics generates 500+ revenue-generating scans daily via the app and WhatsApp.
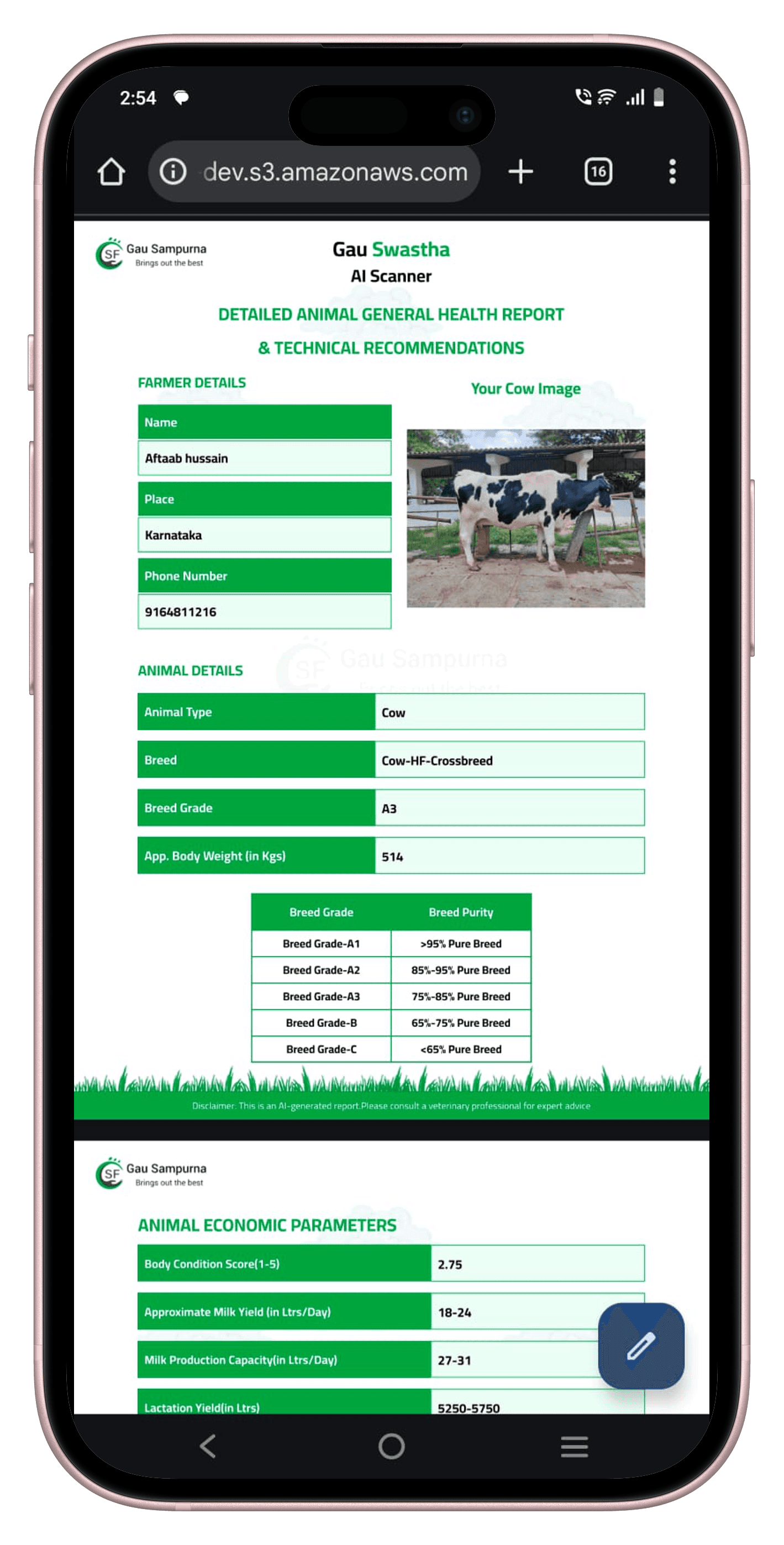
The report covers the metrics that matter most to farmers and vets: breed grade, approximate live weight, body condition score, milk production capacity, lactation stage, and economic parameters. Everything on a single screen, delivered in under two seconds.
Across all 24 classification modules in production, the system averaged 92% top-1 accuracy predicting the right class. Disease detection modules were highest (95%+); BCS scoring was the most challenging due to the fine-grained scale and inherent inter-annotator disagreement — but the three-region ensemble pushed it above the 85% clinical acceptance threshold we set from the start.
GauSampurna on the Google Play Store, grown organically through vet and farmer networks.
Revenue-generating scans through WhatsApp and the app each day — a real commercial signal.
Paid pilot with the Government of Karnataka — real clinical use, not a demo or grant-funded trial.